Snap-shot of current research activities
Our research is focused on developing bioanalytical tools and methodologies and applying them to pertinent and challenging problems in biology and medicine. In developing these bioanalytical tools, we make use of the rich repository of micro- and nanotechnologies that leverage the properties of liquids at the micrometer scale – microfluidics. Microfluidic strategies tend to facilitate new functionalities while allowing for low reagent consumption, precision biochemistry, efficient localization of heat, all the while often are amenable to automation. Our strategies encompass the implemention of quantitative microscale assays that we hope will culminate and provide impetus to the accurate application of personalized medicine.
Open space microfluidics: technologies
Scanning probes for localizing liquids on surfaces
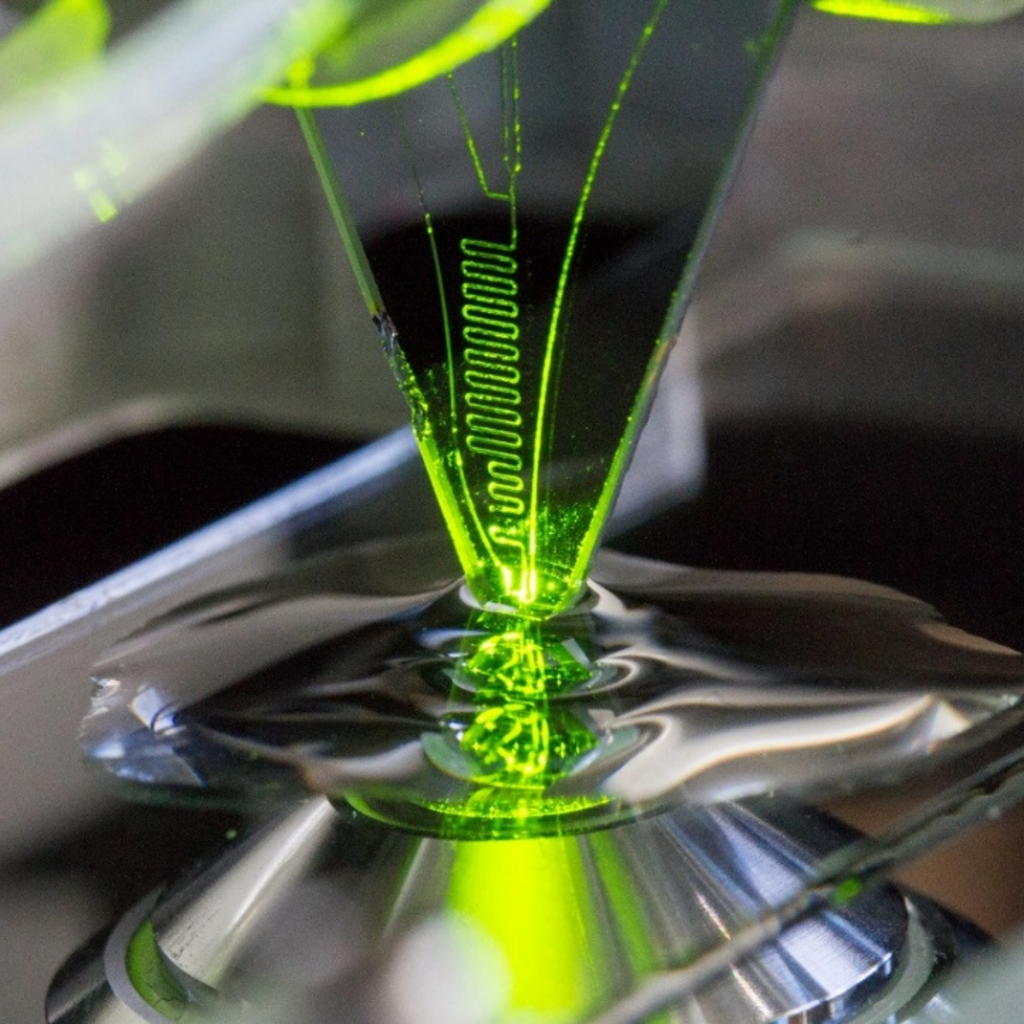

Techniques to study, work and locally probe adherent cells & tissues at micrometer distances from cell surfaces in “open space” would represent a major advance for the biology of biointerfaces. One needs tools for observing, patterning, assaying and stimulating that have a range of dimensions and volumes commensurate with the cell. It is within such situations, new tools and methodologies for discovery and diagnostics play an invaluable role. To this end, we have been advancing several different microfluidics approaches and devices, one of the them being a non-contact, scanning technology, which spatially confines nanoliter volumes of chemicals for interacting with cells at the μm-length scale. This technology is called the vertical microfluidic probe (vMFP) shapes liquid on biological surfaces hydrodynamically (part of this was explored with the framework of an ERC project, BioProbe).
Key references:
Microfluidics in the “Open Space” for Performing Localized Chemistry on Biological Interfaces
G.V. Kaigala, R.D. Lovchik, E. Delamarche
Angewandte Chemie 51(45) 11224-11240, 2012.
A compact and versatile microfluidic probe for local processing of tissue sections and biological specimens
J.F. Cors, R.D. Lovchik, E. Delamarche, G.V. Kaigala
Review of Scientific Instruments 85, 034301, 2014.
Reconfigurable microfluidics: Using fewer number of “walls”
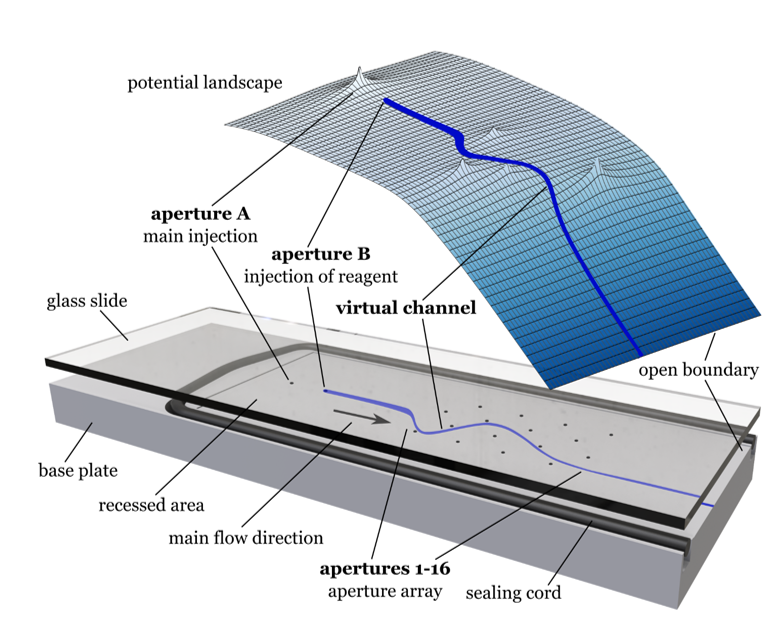
The manipulations of liquids in microscale-sized channels (a.k.a microfluidics) has been applied in the past 30 years to a wide variety of disciplines, with biomedical applications ranging from single cell analysis, analytical tests through tissue section analysis, to creating complex synthetic biological entities. While significant progress has been made in microfluidics, its full potential has not yet been realized, as its use has not fully transcended the developers of the technology to the investigators (users) in the chemical, biological, medical and clinical laboratories. In this project, we introduce a new concept to break the current paradigm in traditional microfluidics that makes use of physical channels and mechanical actuators, in which geometries and functionalities are intimately linked to one another, i.e. changing the flow field
requires change at the mechanical level. We strive to democratize microfluidic devices in order to provide a single platform approach towards addressing the requirements of a wide range of applications. We specifically take two approaches: one is leveraging non-uniform electro-osmotic flows, and the other is using purely hydrodynamics.
Key references:
Reconfigurable Microfluidics: Real-time Shaping of Virtual Channels through Hydrodynamic Forces
Taylor, D., and Kaigala, G. V.
Lab on a Chip, 2020.
Dynamic microscale flow patterning using electrical modulation of zeta potential
Paratore, F., Bacheva, V., Kaigala, G.V., and Bercovici, M.
Proceedings of the National Academy of Sciences, 2019.
Electrokinetics
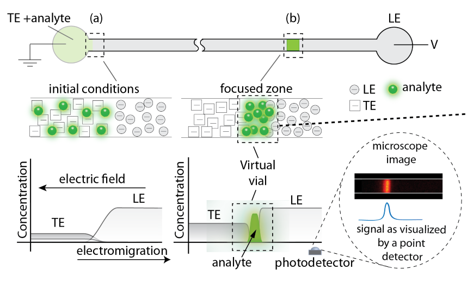
We make use of different eletrokinetic effects in different parts of the project, we are particularly interested in making use of a technique called isoptachophoresis (ITP). ITP allows for simultaneous separation and preconcentration of analytes based on their effective electrophoretic mobility. ITP is very compatible to miniaturized biochemical analysis systems because it is robust to implement, and can achieve high focusing rates in relatively short channels (order 1 cm). A uniting theme of these projects is the use of microfluidic platforms and isotachophoresis (ITP) to create “virtual vials” or “compartments” which enable precise transport, focusing, and control of samples within micron-sized channels.
Key references
- Extraction of electrokinetically separated analytes with on-demand encapsulation
X.F. van Kooten, M. Bercovici, G.V. Kaigala
Lab on a Chip, DOI: 10.1039/C8LC00912K, 2018. - Isotachophoresis-Based Surface Immunoassay
F. Paratore, T. Zeidman Kalman, T. Rosenfeld, G.V. Kaigala, M. Bercovici
Analytical Chemistry 89(14), 7373-7381, 2017. - Focusing analytes from 50uL into 500 pL: On-chip focusing from large sample volumes using isotachophoresis
X.F. van Kooten, M. Truman-Rosentsvit, G.V. Kaigala, M. Bercovici
Scientific Reports 7, 2017.
Leveraging the above bioanalytical tools for selected applications
Surface assays: Biopatterning and Biomolecular responses
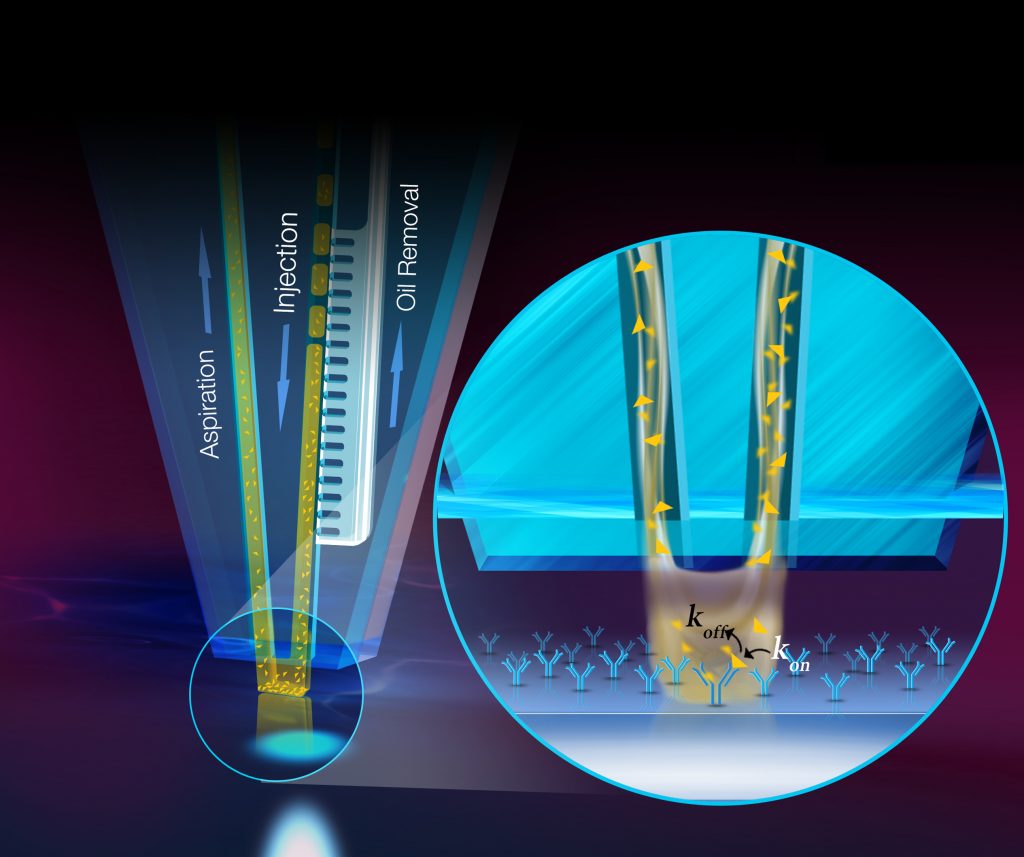
We develop new methodology for efficient and high-quality patterning of biological reagents for surface-based biological assays using tenets of ‘open-space microfluidics’. We study the interplay between diffusion, advection and surface chemistry, and present the design of a non-contact scanning microfluidic device to efficiently present reagents on surfaces. By leveraging convective flows, recirculation and mixing of a processing liquid, this device overcomes limitations of existing biopatterning approaches, such as passive diffusion of analytes, uncontrolled wetting and drying artefacts. We pattern complex protein arrays, DNA arrays and implement antibody-antigen assays on standard biological substrates, and observe high pattern quality, conservative usage of reagents, micrometer-precision of localization and convection-enhanced fast deposition. We think such methods may facilitate quantitative biological assays and spur the development of the next generation of protein microarrays and also help to estimate biomolecular responses.
Key references:
- Underpinning transport phenomena for the patterning of biomolecules
Pereiro, J. F. Cors, S. Pane, B. J. Nelson, and G. V. Kaigala,
Chemical Society Reviews, Vol. 48, Number 5, 2019. - Real-time monitoring of fluorescence in situ hybridization kinetics
Ostromohov, D. Huber, M. Bercovici, and G.V. Kaigala
Analytical Chemistry 90, 11169-11734, 2018. - Convection-enhanced biopatterning with hydrodynamically confined nanoliter volumes of reagents
Autebert, J. Cors, D. Taylor, G.V. Kaigala
Analytical Chemistry 88(6), 3235-3242, 2016.
Tumor profiling systems: Tissue microprocessing for multimodal analysis of tumors
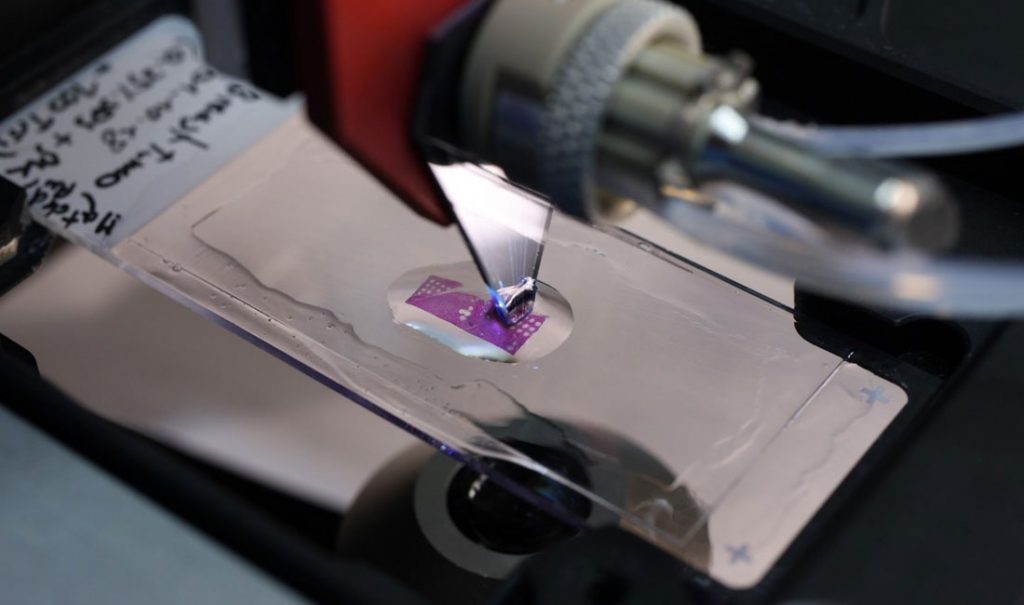

Controlled heterogeneity allows for tissue growth and differentiation in healthy tissues and is a necessary component during tissue morphogenesis. In contrast, uncontrolled heterogeneity is implicated in diseased states of a tissue, where functional compartmentalization of subpopulations is lost. Specifically, heterogeneity in cancer is associated with higher degree of invasiveness of a tumor and often makes therapeutic interventions ineffective. Therefore, a better understanding of the mechanisms that drive tumor progression and other diseased states and their treatment, requires strategies that analyze molecular profiles of biological
samples in the context of heterogeneity. Several of the technologies and methodologies we develop have been designed and directed at interacting with tumor tissues, with an end-goal of enabling more quantitative personalized medicine.
Traditionally, while analyzing tumors for diagnosis, whole biopsies are homogenized prior to downstream processing for profiling. The objective of this work is to develop strategies to perform accurate spatial molecular profiling of heterogeneous tumor tissues and cells in culture by selectively studying cells at multiple length scales. The profiling strategies include multi-modal methods to interrogate genomic, transcriptomic and proteomic characteristics of specific sub-populations within heterogeneous samples. For genomic and transcriptomic studies, a new strategy to selectively lyse cells from heterogeneous ensembles is introduced termed ‘Spatialyse’. For proteomic studies on tumor tissue sections, a new method to quantify biomarker expression using a microscale gradient IHC is introduced termed ‘qµIC – quantitative micro immunohistochemistry’. These methods are being applied to scenarios often observed in cancer research and diagnostics to evaluate their utility.
For more information, please check this page
Key references:
- Quantitative microimmunohistochemistry for the grading of immunostains in tumor tissues,
Kashyap, A. Fomitcheva Khartchenko, P.Pati, M. Gabrani, P.Schraml and G. V. Kaigala,
Nature Biomedical Engineering, 3, 478–490, 2019. - Selective local lysis and sampling of live cells for nucleic acid analysis using a microfluidic probe
Kashyap, J. Autebert, E. Delamarche, G.V. Kaigala
Scientific Reports 6, 29579, 2016. - Micro fluorescence in situ hybridization (µFISH) for spatially multiplexed analysis of a cell monolayer
Huber, J. Autebert, G.V. Kaigala
Biomedical Microdevices 18(40), 2016. - High-Quality Immunohistochemical Stains through Computational Assay Parameter Optimization
Arar, N. M. ; Pati, P., Kashyap, P. ; Fomitcheva Khartchenko, A., Goksel, O., Kaigala, G. V., Gabrani, M.
IEEE Transaction on Biomedical Engineering, 2019.
